R.D. Pascual-Marqui
The KEY Institute for Brain-Mind Research, University Hospital of Psychiatry
Lenggstr. 31, CH-8029 Zurich, Switzerland
Tel.:+41-1-3884934 ; Fax:+41-1-3803043
pascualm@key.unizh.ch ; http://www.keyinst.unizh.ch/loreta.htm
Scalp electric potentials (EEG) and extracranial magnetic fields (MEG) are due to the primary (impressed) current density distribution that arises from neuronal post-synaptic processes. A solution to the inverse problem, i.e. the computation of images of electric neuronal activity based on extracranial measurements, would provide important information on the time course and localization of brain function. In general, there is no unique solution to this problem. In particular, an instantaneous, distributed, discrete, linear solution capable of exact localization of point sources is of great interest, since the principles of linearity and superposition would guarantee its trustworthiness as a functional imaging method, given that brain activity occurs in the form of a finite number of distributed “hot spots”. Despite all previous efforts, linear solutions at best produced images with systematic non-zero localization errors. A solution is reported here, which yields images of standardized current density with zero localization error. The purpose of this paper is to present the technical details of the method, allowing researchers to test, check, reproduce, and validate the new method (sLORETA).
This study is strictly
limited to EEG/MEG inverse solutions of the type: instantaneous, distributed,
discrete, and linear. The generic form of the inverse problem follows. There
are ![]() instantaneous
extracranial measurements. There are
instantaneous
extracranial measurements. There are ![]() voxels in the brain. Typically, the voxels
are determined by subdividing uniformly the solution space, which is usually
taken as the cortical grey matter volume or surface. At each voxel there is
a point source, which may be a vector with three unknown components (i.e., the
three dipole moments), or a scalar (unknown dipole amplitude, known orientation).
The cases considered here correspond to
voxels in the brain. Typically, the voxels
are determined by subdividing uniformly the solution space, which is usually
taken as the cortical grey matter volume or surface. At each voxel there is
a point source, which may be a vector with three unknown components (i.e., the
three dipole moments), or a scalar (unknown dipole amplitude, known orientation).
The cases considered here correspond to ![]() .
.
In 1984, Hämäläinen and Ilmoniemi (1) were the first to report an instantaneous, distributed, discrete, linear solution to the EEG/MEG inverse problem: the well known minimum norm solution. However, the minimum norm solution is notorious for totally misplacing actual deep sources onto the outermost cortex, as demonstrated in (3), (2), and (8).
The problem of excessively large errors of localization remained unsolved until the introduction of the method known as LORETA (low resolution brain electromagnetic tomography) in 1994 (18). LORETA has fairly good accuracy in localizing test sources even when they are deep. The overall average localization error is smaller than one grid unit (see e.g. (3), (2), and (8)).
A series of papers published in 1998 and 1999 (see (19)-(23)) introduced for the first time the method of high time resolution statistical parametric mapping for tomographic images of electric neuronal activity. The idea was to adopt the methods of statistical inference for the localization of brain function as used in PET and fMRI studies. This methodology was applied to high time resolution, time varying LORETA images.
In the present study a new tomographic method for electric neuronal activity is introduced, where localization inference is based on images of standardized current density. The method is denoted as standardized low resolution brain electromagnetic tomography (sLORETA). Unlike the method recently introduced by Dale et al. (6), which has systematic non-zero localization error, sLORETA has zero localization error.
The equation of interest takes the form:
![]() Eq. 1
Eq. 1
In Eq. 1, ![]() is a vector containing
scalp electric potentials measured at
is a vector containing
scalp electric potentials measured at ![]() cephalic electrodes, with respect to
a common, arbitrary reference electrode located anywhere on the body.
cephalic electrodes, with respect to
a common, arbitrary reference electrode located anywhere on the body.
The primary (impressed)
current density ![]() is defined as:
is defined as:
![]() Eq. 2
Eq. 2
where ![]() for
for ![]() . At the lth voxel,
. At the lth voxel, ![]() contains the three unknown
dipole moments.
contains the three unknown
dipole moments.
The superscript “T” denotes transpose.
The lead field ![]() has the following
structure:
has the following
structure:
 Eq. 3
Eq. 3
with ![]() , for
, for ![]() , and for
, and for ![]() . Note that
. Note that ![]() , where
, where ![]() is the scalp electric
potential at the ith
electrode, due to a unit strength X-oriented
dipole at the lth voxel;
is the scalp electric
potential at the ith
electrode, due to a unit strength X-oriented
dipole at the lth voxel;
![]() is the
scalp electric potential at the ith
electrode, due to a unit strength Y-
oriented dipole at the lth
voxel; and
is the
scalp electric potential at the ith
electrode, due to a unit strength Y-
oriented dipole at the lth
voxel; and ![]() is
the scalp electric potential at the ith
electrode, due to a unit strength Z-
oriented dipole at the lth
voxel.
is
the scalp electric potential at the ith
electrode, due to a unit strength Z-
oriented dipole at the lth
voxel.
In Eq. 1, c is an arbitrary constant which embodies
the fact that the electric potential is determined up to an arbitrary constant;
and ![]() is
a vector of ones. The parameter c
allows the use of any reference for the lead field and the measurements.
is
a vector of ones. The parameter c
allows the use of any reference for the lead field and the measurements.
Hämäläinen, M.S., and Ilmoniemi (1) were the first to publish a particular solution to the instantaneous, distributed, discrete, linear EEG/MEG inverse problem. Their solution is known as the minimum norm inverse solution. However, the minimum norm solution is notorious for totally misplacing actual deep sources onto the outermost cortex (2).
Dale et al. (6) proposed a method in which localization inference is based on a standardization of the current density estimates. In particular, they employed the current density estimate given by the minimum norm solution, and they standardized it by using its expected standard deviation, which is hypothesized to be originated exclusively by measurement noise. The method of Dale et al. (6) produces systematic non-zero localization errors (8), even in the presence of negligible noise. This fact was not evaluated nor admitted in their original paper.
sLORETA is similar to the Dale et al. (6) method: it employs the current density estimate given by the minimum norm solution, and localization inference is based on standardized values of the current density estimates. However, standardization in sLORETA takes a completely different route (explained below). The consequence is that, unlike the Dale et al. (6) method, sLORETA is capable of exact (zero-error) localization.
The minimum norm inverse
solution is harmonic (2), which means that the Laplacian of the current density
is zero, i.e., ![]() , where r denotes volume coordinates in the brain. Therefore, the minimum
norm inverse solution is very smooth. The concept of smoothness employed here
is discussed in greater detail in (8), with special emphasis on its electrophysiological
interpretation. However, as previously mentioned, the minimum norm inverse solution
is notorious for its incapability of correct localization of deep point sources
(2).
, where r denotes volume coordinates in the brain. Therefore, the minimum
norm inverse solution is very smooth. The concept of smoothness employed here
is discussed in greater detail in (8), with special emphasis on its electrophysiological
interpretation. However, as previously mentioned, the minimum norm inverse solution
is notorious for its incapability of correct localization of deep point sources
(2).
This problem is solved by standardization of the minimum norm inverse solution, and basing localization inference on these standardized estimates.
The functional of interest here is:
![]() Eq. 4
Eq. 4
where ![]() is a regularization
parameter. This functional is to be minimized with respect to J and c,
for given K, F,
and a.
The explicit solution to this minimization problem is (see e.g. (3)):
is a regularization
parameter. This functional is to be minimized with respect to J and c,
for given K, F,
and a.
The explicit solution to this minimization problem is (see e.g. (3)):
![]() Eq. 5
Eq. 5
where:
![]() Eq. 6
Eq. 6
![]() Eq. 7
Eq. 7
with ![]() denoting the centering
matrix;
denoting the centering
matrix; ![]() the
identity matrix; and
the
identity matrix; and ![]() is a vector of ones.
is a vector of ones.
For any matrix ![]() ,
, ![]() denotes its Moore-Penrose
pseudoinverse (see e.g. (9)).
denotes its Moore-Penrose
pseudoinverse (see e.g. (9)).
The centering matrix H in Eq. 7 is the average reference operator.
In what follows, for all EEG cases, the symbols F and K will denote the average reference transforms of the EEG measurements and the lead field, respectively. This simplifies the notation. But most important of all, the correct solution to EEG problems is based on these average reference transforms.
Therefore, when using average reference transforms of F and K, Eq. 1 becomes:
![]() Eq. 8
Eq. 8
and the functional in Eq. 4 becomes:
![]() Eq. 9
Eq. 9
with minimum:
![]() Eq. 10
Eq. 10
where:
![]() Eq. 11
Eq. 11
Standardization of the
estimate ![]() requires
an estimate of its variance.
requires
an estimate of its variance.
Note that Eq. 9 can be
derived from a Bayesian formulation of the inverse problem (see e.g. (5), Eq.
1.88 therein). In this view, the actual
source variance (prior) ![]() is equal to the identity
matrix, i.e.:
is equal to the identity
matrix, i.e.:
![]() Eq. 12
Eq. 12
In addition, from the Bayesian point of view, the electric potential variance is due to noisy measurements:
![]() Eq. 13
Eq. 13
Note that in Eq. 13, the average reference operator H plays the role of the identity matrix in the subspace spanned by the measurement space.
It is usually assumed that activity of the actual sources and the noise in the measurements are uncorrelated.
Based on the linear relation
in Eq. 8, making use of Eqs. 12 and 13, and taking into account the independence
of actual source activity and measurement noise, the electric potential variance
![]() then is:
then is:
![]() Eq. 14
Eq. 14
See e.g. (10), Eqs. 1.5.1-1.5.6 therein.
Due to the linear relation in Eq. 10, and making use of Eq. 14, the variance of the estimated current density is:
![]() Eq. 15
Eq. 15
See e.g. (10), Eqs. 1.5.1-1.5.6 therein, and (9).
Note that the variance of the estimated current density is equivalent to the Backus and Gilbert (4) resolution matrix, which is obtained by plugging Eqs. 8 and 11 into 10:
![]() Eq. 16
Eq. 16
with:
![]() Eq. 17
Eq. 17
where R is the resolution matrix.
Note that the variance of the estimated current density in Eqs. 15 and 17 is not the posterior variance in the Bayesian formulation (see e.g. (5), Eq. 1.94).
In contrast, according to Dale et al. (6), the variance of the estimated current density is based on the assumption that the only source of variation is measurement noise. This means that Eq. 14 now is:
![]() Eq. 18
Eq. 18
and Eq. 15 now is:
![]() Eq. 19
Eq. 19
Note that unlike the approach of Dale et al. (6), sLORETA takes into account two sources of variation: mainly the variation of the actual sources, and then finally, if any, the variation due to noisy measurements.
Finally, sLORETA corresponds to the following estimates of standardized current density power:
![]() Eq. 20
Eq. 20
where ![]() is the current density
estimate at the lth
voxel given by Eqs. 10 and 11 (for average reference transforms); and
is the current density
estimate at the lth
voxel given by Eqs. 10 and 11 (for average reference transforms); and ![]() is the lth diagonal block of matrix
is the lth diagonal block of matrix ![]() in Eqs. 15 or 17.
in Eqs. 15 or 17.
Note that the pseudo-statistic in Eq. 20 has the form of an “F” statistic.
Note that Eq. 20 is different in form from the Dale et al. (6) standardization (see (6), Eq. 7 therein). The Dale et al. (6) standardized estimates are:
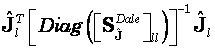 Eq. 21
Eq. 21
where ![]() is the lth diagonal block of matrix
is the lth diagonal block of matrix ![]() in Eq. 19; and for
any symmetric matrix M,
in Eq. 19; and for
any symmetric matrix M, ![]() is the diagonal matrix
formed by the diagonal elements of M.
is the diagonal matrix
formed by the diagonal elements of M.
This case usually corresponds to the inverse problem when the cortical surface is completely known. Voxels are now distributed along the cortical surface, and the dipoles at each voxel have known orientation (perpendicular to the cortical surface). The unknowns correspond to the amplitudes, which may take positive, zero, or negative values. The dipole orientations (defined as unit length vectors with three components) can be incorporated into the lead field K in Eq. 8. Details can be found in (3).
In this case Eq. 8 has
the same form, but now ![]() since it only contains one unknown scalar
per voxel, and
since it only contains one unknown scalar
per voxel, and ![]() since it includes the dipole orientation
at each voxel. Details can be found in (3). All the derivations employed in
Eqs. 8-17 remain formally identical.
since it includes the dipole orientation
at each voxel. Details can be found in (3). All the derivations employed in
Eqs. 8-17 remain formally identical.
However, sLORETA now corresponds to the following estimates of standardized current density power:
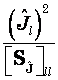 Eq. 22
Eq. 22
where the scalar
![]() is the
current density amplitude estimate at the lth
voxel; and the scalar
is the
current density amplitude estimate at the lth
voxel; and the scalar ![]() is the lth diagonal element of matrix
is the lth diagonal element of matrix ![]() .
.
Note that the pseudo-statistic in Eq. 22 has the form of an “F” statistic.
The equations for the MEG case have identical form to Eqs. 8-17, 20, and 22, depending on the case of unknown dipole moments, or only unknown amplitudes.
Note that the average reference does not apply to MEG.
The only change corresponds to the equations for the MEG lead field, which are different to those for the EEG.
Simulations were carried out in a three-shell spherical head model registered to the Talairach human brain atlas (11), available as a digitized MRI from the Brain Imaging Centre, Montreal Neurological Institute. Registration between spherical and realistic head geometry used EEG electrode coordinates reported by Towle et al. (12).
In one set of practical, realistic, simulations, the solution space was restricted to cortical gray matter and hippocampus, as determined by the corresponding digitized Probability Atlas also available from the Brain Imaging Centre, Montreal Neurological Institute. A voxel was labeled as gray matter if it met the following three conditions: its probability of being gray matter was higher than that of being white matter, its probability of being gray matter was higher than that of being cerebrospinal fluid, and its probability of being gray matter was higher than 33%. Only gray matter voxels that belonged to cortical and hippocampal regions were used for the analysis. A total of 6430 voxels at 5mm spatial resolution were produced under these neuroanatomical constraints. At each voxel, three unknown values (the three dipole moments) were estimated, making a total of 6430x3=19290 unknowns. 25 electrodes (in EEG experiments), or 25 magnetometer sensors (in MEG experiments) were used. In both cases, sensors and electrodes were placed in the same locations.
In the second set of practical, realistic, simulations, the solution space was restricted to the cortical surface, represented as 12980 triangles (voxels) (13). This case corresponded to unknown current density amplitude (but with known orientation), making a total of 12980 unknowns. 101 electrodes (in EEG experiments), or 101 magnetometer sensors (in MEG experiments) were used. In both cases, sensors and electrodes were placed in the same locations.
The minimum norm solution, the method of Dale et al. (6), and sLORETA were compared in terms of localization errors and spatial spread. The methods were tested with point sources located at the voxels. For the case corresponding to 3 unknowns per voxel, an arbitrary (random) orientation of the test source was employed. The test sources were used to generate the measurements (forward equation (Eq. 8)), which were then given to the imaging methods. Simulations included “noise free” and “noisy” measurements.
In the minimum norm solution
case, the imaging method is based on Eqs. 20 and 22, but without standardization,
which is achieved by setting the variance to the identity matrix, i.e., ![]() .
.
In the minimum norm solution and in sLORETA, the regularization parameter a in the previous equations was estimated by cross-validation. Exact details and equations for a practical implementation of the cross-validation method can be found in (14).
In the Dale et al. (6) method, the parameter a is interpreted as the variance of the noise in the measurements, and this value was determined by the simulation design. In the “noise free” case, a very small value of a was used, typically in the order of 10-10 times the power of the scalp field produced by the test source with lowest scalp field power.
Localization error was defined as the distance between the actual test source and the location of the maximum in the imaging method. The spatial spread was defined identically as in (3), which corresponds to a measure of spatial standard deviation of the imaging method centered at the actual test source, and not at the imaging method’s own maximum, since this would unjustifiably favor the method’s performance.
Figures 1a-1h summarize localization error, spatial spread, and estimated activity values for the three imaging methods (minimum norm, Dale, and sLORETA).
Note that the estimated activity values at test source locations cannot be compared among the different imaging methods, since these values are in different units for the different imaging methods. However, this feature is very informative for comparing the quality of the different methods. For example, from Fig 1a, the ratios of estimated source activity (maximum to minimum) were 850, 103, and, 30, for minimum norm, Dale, and sLORETA, respectively. This means that with sLORETA, some sources (especially deep ones) will be underestimated. However, sLORETA out-performs tremendously the minimum norm and the Dale methods in this aspect.
In all noise free simulations, only sLORETA has exact, zero error localization. In all noisy simulations, sLORETA has by far the lowest localization errors. In most cases, the spatial spread (i.e. “blurring”) of sLORETA is smaller than that of the Dale method.
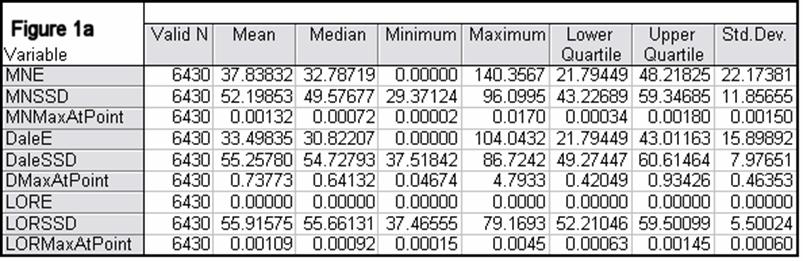
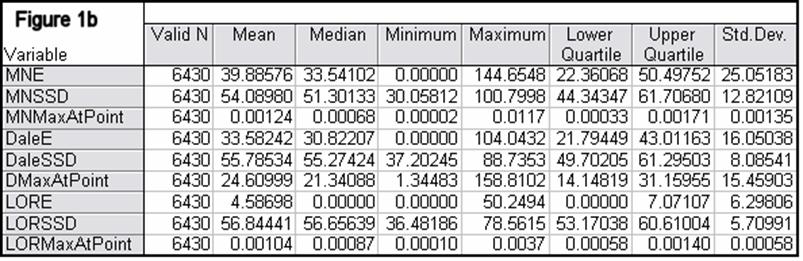
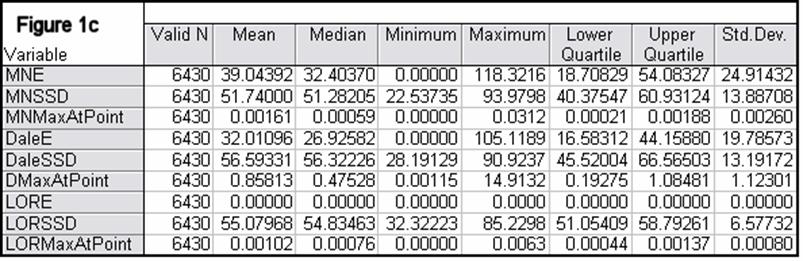
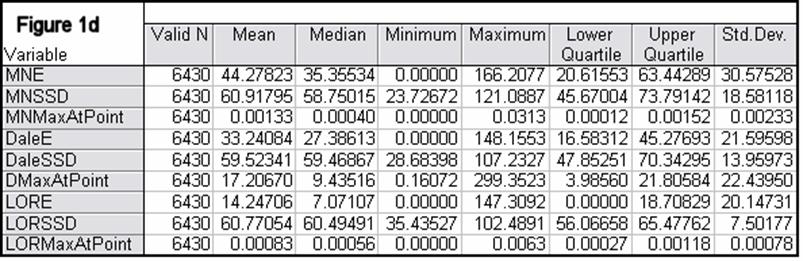
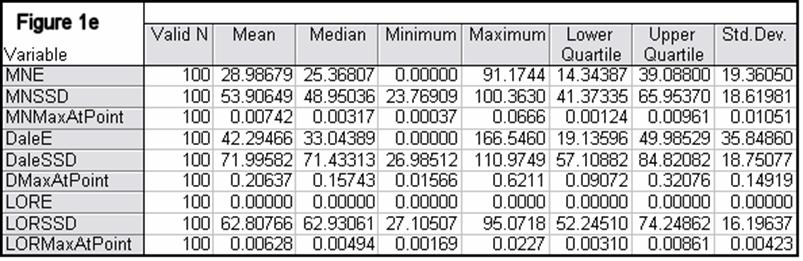
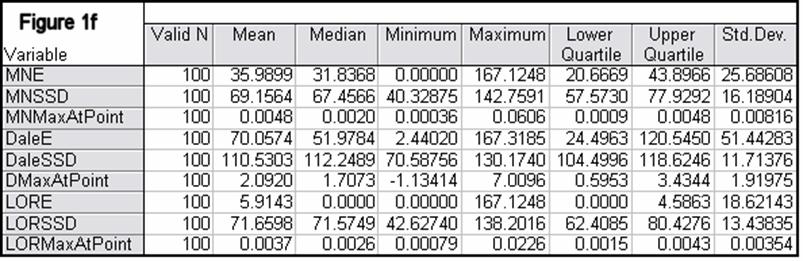
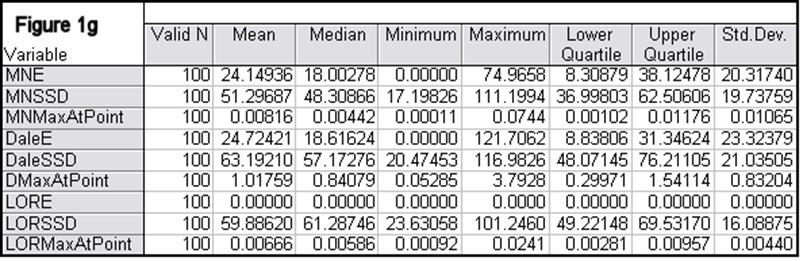
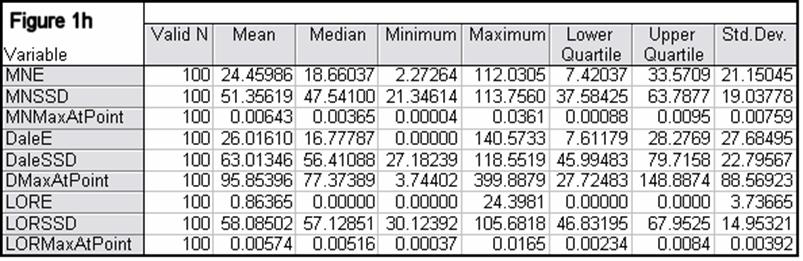
Figure 1: “a-h” summarize in tabular form localization error, spatial spread, and estimated activity values for the three imaging methods (minimum norm, Dale, and sLORETA). (a) EEG, 6430 voxels, 3 unknowns per voxel, 25 electrodes, 6430 test sources with random orientation, no noise. (b) Same as (a), but with additive random noise (noise scalp field standard deviation equal to 0.12 times the test source with lowest scalp field standard deviation). (c) MEG, 6430 voxels, 3 unknowns per voxel, 25 sensors, 6430 test sources with random orientation, no noise. (d) Same as (c), but with additive random noise (noise scalp field standard deviation equal to 7.21 times the test source with lowest scalp field standard deviation). (e) EEG, 12980 voxels, 1 unknown per voxel, 101 electrodes, 100 randomly selected test sources, no noise. (f) Same as (e), but with additive random noise (noise scalp field standard deviation equal to 0.082 times the test source with lowest scalp field standard deviation). (g) MEG, 12980 voxels, 1 unknown per voxel, 101 sensors, 100 randomly selected test sources, no noise. (h) Same as (g), but with additive random noise (noise scalp field standard deviation equal to 8.49 times the test source with lowest scalp field standard deviation). MNE: minimum norm localization error (mm); MNSSD: minimum norm spatial standard deviation (mm); MNMaxAbs: estimated minimum norm activity value at test source location (arbitrary units); DaleE: Dale localization error (mm); DaleSSD: Dale spatial standard deviation (mm); DMaxAbs: estimated Dale activity value at test source location (arbitrary units); LORE: sLORETA localization error (mm); LORSSD: sLORETA spatial standard deviation (mm); LORMaxAbs: estimated sLORETA activity value at test source location (arbitrary units). Note that the estimated activity values at test source locations cannot be compared among the different imaging methods (see text for explanation).
The main properties of sLORETA, for both EEG and MEG, based on estimates of activity given by Eq. 20 are:
1. Exact, zero error, localization for test dipoles located at voxel positions, in the absence of noisy measurements.
2. Exact, zero error, localization of test dipoles with arbitrary orientation, located at voxel positions, in the absence of noisy measurements.
3. Exact, zero
error, localization of test dipoles with
arbitrary orientation, located at voxel positions, in the absence
of noisy measurements, even under regularization ![]() .
.
4. Exact, zero error, localization even for dipoles corresponding to a non-connected grid. For example, cortical and non-connected subcortical grey matter can now be modeled as the solution space. The error remains zero.
The main properties of sLORETA based on estimates of activity given by Eq. 22 are:
1. Exact, zero error, localization of test dipoles located at voxel positions, in the absence of noisy measurements.
3. Exact, zero
error, localization of test dipoles located at voxel positions, in the absence
of noisy measurements, even under regularization ![]() .
.
4. Exact, zero error, localization even for dipoles corresponding to non-connected grids. For example, cortical and non-connected subcortical grey matter can now be modeled as the solution space. The error remains zero.
5. These results mean that the distribution of voxels can be quite arbitrary. For example, voxels do not have to be uniformly distributed from the geometrical point of view, although they should be uniformly distributed from the “grey matter density” point of view. Furthermore, different types of voxels may exist, some with unknown current density vector, and some with known current density orientation but unknown amplitude.
Suppose there exist reasons to believe that the actual (prior) current density variance is the diagonal, positive definite matrix W. This situation arises for example, in some approaches that force fMRI hot spot locations onto the EEG/MEG inverse solution (see for example (6)). In this case, Eq. 8 can be rewritten as:
![]() Eq. 23
Eq. 23
where the new
unknown variable ![]() has
been “pre-standardized” to have the identity matrix as its variance. This transformed
variable plays the role of J in
all equations above, and
has
been “pre-standardized” to have the identity matrix as its variance. This transformed
variable plays the role of J in
all equations above, and ![]() plays the role of the lead field in all
equations above. All else proceeds identically with these new formal substitutions.
plays the role of the lead field in all
equations above. All else proceeds identically with these new formal substitutions.
Note that the final sLORETA image corresponds to standardized
estimates of activity (Eqs. 20 or 22) for the pre-standardized current density
![]() .
.
Note that this approach can be applied to any actual (prior) current density variance W, as long as it is positive definite, and there exists a meaningful decomposition:
![]() Eq. 24
Eq. 24
for the square
root matrix ![]() .
For example, this is the case of the classical LORETA method (2), where
.
For example, this is the case of the classical LORETA method (2), where ![]() embodies a discrete
spatial Laplacian operator that achieves smoothness between neighboring voxels.
embodies a discrete
spatial Laplacian operator that achieves smoothness between neighboring voxels.
The regularization parameter a cannot be estimated from the functional in Eq. 8. However, it can be estimated via the cross-validation functional. This has been published in (14), in a reply to comments made to the paper in (2), which includes the detailed derivation of the method, and a set of equations that can be used efficiently in practice.
Although sLORETA calculations produce pseudo-statistics, it is highly recommended to not use these values as actual statistics in testing of hypotheses in experimental designs.
Unlike the approach of Dale et al. (6), which makes use of their statistics for hypothesis testing, it is recommended to use sLORETA pseudo-statistic values as estimates of activity, and to apply standard techniques such as in statistical non-parametric mapping (SnPM) (7) for the analysis of experimental designs.
Note that tests for absolute activation with sLORETA can be performed by using the modified pseudo-statistics:
![]() Eq. 25
Eq. 25
or:
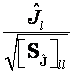 Eq. 26
Eq. 26
which correspond to Eqs. 20 and 22, respectively.
These pseudo-statistics
should be used in an experimental design where there are N independent sLORETA images. For example, in a visual event related potential
study with ![]() subjects, consider the 10 sLORETA images at the P100 latency.
subjects, consider the 10 sLORETA images at the P100 latency.
In Eq. 25, ![]() denotes the unique
symmetric inverse square root matrix of
denotes the unique
symmetric inverse square root matrix of ![]() . The pseudo-statistic in Eq. 26 has the
form of a univariate Student’s t-statistic,
and the pseudo-statistic in Eq. 25 has the form of a Mahalanobis transform (10).
. The pseudo-statistic in Eq. 26 has the
form of a univariate Student’s t-statistic,
and the pseudo-statistic in Eq. 25 has the form of a Mahalanobis transform (10).
In the case of unknown amplitudes only, significant absolute activation is based on testing for zero mean of the pseudo-statistic in Eq. 26. SnPM can be used to correct for multiple comparisons and to bypass assumptions of Gaussianity.
In the case of unknown current density vector, significant absolute activation is based on testing for zero mean of the “max-statistic” of the pseudo-statistic in Eq. 25. This corresponds to the maximum of the absolute value among the three components. The “max-statistic” reduces three numbers per voxel to a single number per voxel. This is then used in SnPM to correct for multiple comparisons and to bypass assumptions of Gaussianity.
1. Localization error can not be improved beyond the present result. It is zero. Up to the present, no other instantaneous, distributed, discrete, imaging method for EEG/MEG has been published (to the best of the author’s knowledge) that achieved perfect localization. All other previously published methods at best produced systematic non-zero localization errors (see (2), (6), (15), (16), (17)).
2. If the aim is localization, this new method, denoted as sLORETA, at least has perfect first order localization.
3. A distributed imaging method capable of exact localization of point sources is of great interest, since the principles of linearity and superposition would guarantee its trustworthiness as a functional imaging method, given that brain activity occurs in the form of a finite number of distributed “hot spots”.
4. The detailed information provided here allows the reader to reproduce, check, test, and validate the previous claims.
1) Hämäläinen, M.S., and Ilmoniemi, R.J. Interpreting measured magnetic fields of the brain: estimates of current distributions. Tech. Rep. TKK-F-A559, Helsinki University of Technology, Espoo, 1984.
2) Pascual-Marqui RD. Review of methods for solving the EEG inverse problem. International Journal of Bioelectromagnetism 1999, 1: 75-86.
3) Pascual-Marqui RD. Reply to comments by Hämäläinen, Ilmoniemi and Nunez. In Source Localization: Continuing Discussion of the Inverse Problem (W. Skrandies, Ed.), ISBET Newsletter, 1995, No. 6, pp:16-28 (ISSN 0947-5133).
4) Backus G and Gilbert F. The resolving power of gross earth data. Geophys. J. R. Astr. Soc. 1968, 16:169-205.
5) A. Tarantola. Inverse problem theory: methods for data fitting and model parameter estimation. Elsevier, Amsterdam, 1987.
6) Dale AM, Liu AK, Fischl BR, Buckner RL, Belliveau JW, Lewine JD, Halgren E. Dynamic statistical parametric mapping: combining fMRI and MEG for high-resolution imaging of cortical activity. Neuron 2000, 26: 55-67.
7) Nichols TE, Holmes AP. Nonparametric permutation tests for functional neuroimaging: a primer with examples. Human Brain Mapping 2001, 15: 1–25.
8) Pascual-Marqui RD, Esslen M, Kochi K, Lehmann D. Functional imaging with low resolution brain electromagnetic tomography (LORETA): review, new comparisons, and new validation. Japanese Journal of Clinical Neurophysiology 2002, 30:81-94.
9) C.R. Rao and S.K. Mitra. Theory and application of constrained inverse of matrices. SIAM J. Appl. Math., 1973, 24: 473-488.
10) K.V. Mardia, J.T. Kent, and J.M. Bibby. Multivariate Analysis. Academic Press, London, 1979.
11) Talairach J and Tournoux P. Co-Planar Stereotaxic Atlas of the Human Brain. Thieme, Stuttgart. 1988.
12) Towle VL, Bolanos J, Suarez D, Tan K, Grzeszczuk R, Levin DN, Cakmur R, Frank SA, and Spire JP. The spatial location of EEG electrodes: locating the best-fitting sphere relative to cortical anatomy. Electroencephalography and Clinical Neurophysiology 1993, 86, 1-6.
13) Dickson J, Drury H., Van Essen DC. ‘The surface management system’ (SuMS) database: a surface-based database to aid cortical surface reconstruction, visualization and analysis. Philosophical Transactions of the Royal Society, London, B 2001, 356: 1277-1292. Cortices downloadable at: http://stp.wustl.edu
14) Pascual-Marqui RD. Reply to Comments Made by R. Grave De Peralta Menendez and S.I. Gozalez Andino. International Journal of Bioelectromagnetism 1999, Vol. 1, No. 2, at: http://www.ee.tut.fi/rgi/ijbem/volume1/number2/html/pascual.htm. [View HTML or PDF]
15) Menendez RGD, Andino SG, Lantz G, Michel CM, Landis T. Noninvasive localization of electromagnetic epileptic activity. I. Method descriptions and simulations. Brain Topography. 2001, 14:131-137.
16) Phillips C, Rugg MD, Friston KJ. Systematic regularization of linear inverse solutions of the EEG source localization problem. Neuroimage. 2002, 17:287-301.
17) Phillips C, Rugg MD, Friston KJ. Anatomically informed basis functions for EEG source localization: Combining functional and anatomical constraints. Neuroimage. 2002, 16:678-695.
18) Pascual-Marqui RD, Michel CM, Lehmann D. Low resolution electromagnetic tomography: a new method for localizing electrical activity in the brain. International Journal of Psychophysiology. 1994, 18:49-65.
19) Strik WK, Fallgatter AJ, Brandeis D, Pascual-Marqui RD. Three-dimensional tomography of event-related potentials during response inhibition: evidence for phasic frontal lobe activation. Evoked Potentials-Electroencephalography and Clinical Neurophysiology 1998, 108:406-413.
20) Anderer P, Pascual-Marqui RD, Semlitsch HV, Saletu B. Electrical sources of P300 event-related brain potentials revealed by low resolution electromagnetic tomography .1. Effects of normal aging. Neuropsychobiology 1998, 37:20-27.
21) Anderer P, Saletu B, Semlitsch HV, Pascual-Marqui RD. Electrical sources of P300 event-related brain potentials revealed by low resolution electromagnetic tomography .2. Effects of nootropic therapy in age-associated memory impairment. Neuropsychobiology 1998, 37:28-35.
22) Anderer P, Pascual-Marqui RD, Semlitsch HV, Saletu B. Differential effects of normal aging on sources of standard N1, target N1 and target P300 auditory event-related brain potentials revealed by low resolution electromagnetic tomography (LORETA). Evoked Potentials-Electroencephalography and Clinical Neurophysiology 1998, 108:160-174.
23) Pascual-Marqui RD, Lehmann D, Koenig T, Kochi K, Merlo MCG, Hell D, Koukkou M. Low resolution brain electromagnetic tomography (LORETA) functional imaging in acute, neuroleptic-naive, first-episode, productive schizophrenia. Psychiatry Research-Neuroimaging 1999, 90:169-179.